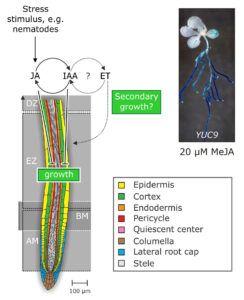
Figure 4. Working-model of the physiological mode of action of the crosstalk between jasmonate, auxin, and ethylene in the framework of stress responses of the root.
CURRENT RESEARCH PROJECTS OF THE POLLMANN LAB - Plant hormone crosstalk
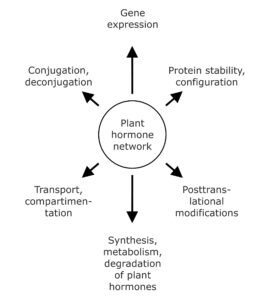
Over recent years, mounting evidence led to the widely accepted concept that plant hormone action is not the read-out of linear pathways, but determined by the extensive combinatorial activity of the signaling molecules and the integration of their signaling pathways, both in terms of regulating growth and development and in adapting to external stimuli. Further complicating the scenario, the triggered physiological processes are not only dependent on the perceived stimulus, but also on the specific properties of the responding tissue in terms of sensitivity and responsiveness to a given signaling molecule class. The plant hormone network can affect plant development and physiological responses on several different levels involving for example, control of mRNA and protein synthesis, configuration, modification, and turnover of proteins, as well as by hormone transport and reversible or irreversible inactivation of active signaling molecules (Fig. 1). For instance, directed polar auxin transport and spatiotemporally defined auxin maxima, and the therewith coupled control of gene expression and gene signaling hierarchies, coordinate organogenesis and axis formation during embryogenesis and patrol plant development.
Plant hormone action largely depends on the combinatorial interplay of the plant hormones, rather than on the individual activity of isolated phytohormones
Another example is the modification of plant hormones by, e.g., glycosylation, methylation, or amino acid conjugation, which either serve to reversibly modulate their activity or confer the first step in their irreversible metabolism. In addition, plant hormones can also directly impact the synthesis or degradation of other signaling molecules, as has been shown, amongst other relationships, for ethylene production, which is induced both by auxin and brassinosteroids. Also, posttranslational modifications of proteins can be a target modulated by plant hormones, as is the case in cytokinin, abscisic acid, and ethylene signaling that involve phosphorylation of downstream components of the signal transduction machinery.
Figure 1. The plant hormone network affects plant development on several different levels. A plethora of different physiological and biochemical processes constitute possible targets of phytohormone action. They reach from the regulation of gene expression and protein abundance over posttranslational modifications of proteins and the influence of plant hormone homeostasis by affecting synthesis, metabolism, and conjugation of signaling compounds to the impact on hormone transport and compartmentation.
Investigation of hormonal crosstalk is challenging, as for a long time hormone functions have largely been studied separately, which makes operational data integration difficult to achieve. For example, it wasn’t until the beginning of this century that the plant scientific community was able to answer the need for simultaneous plant hormone analysis technologies (Müller et al. 2002; Chiwocha et al. 2003; Schmelz et al. 2003; Durgbanshi et al. 2005), which was, in fact for quite some time largely hampered by inadequate analytical means. Nowadays, the available technologies, whether it is transcript profiling at a whole-genome scale, or non-targeted metabolite profiling by cutting-edge mass spectrometric techniques, have greatly improved. Thus, major breakthroughs in deciphering interaction nodes within the hormonal network and in resolving the intricate regulatory patterns in this framework may be expected within the next five to ten years. This will likely include discovery of distinct responses of the different signaling compounds to defined internal and external stimuli, and how these signals are integrated on the molecular level. In order to achieve this goal, a profound understanding of feedback and feed-forward loops, as well as of spatiotemporal regulation of hormone contents at the cellular, tissue, and organ level is required.
In this context, the aim of our lab is to shed light on the mechanistic and conceptual crosstalk between auxin and oxylipins, in particular between indole-3-acetic acid and jasmonic acid; two plant hormones that, at first glance, appear to have not much in common. In previous studies, taking an microarray approach, we identified two YUCCA genes, YUC8 and YUC9, to respond differentially to a treatment with methyl jasmonate (Fig. 2).
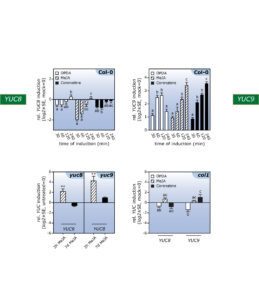
Figure 2. Genome-wide microarray and qPCR analyses provided evidence for the differential impact of oxylipins on YUC8 and YUC9 gene expression.
Further experiments confirmed the effect of methyl jasmonate on the expression of the two YUCCA genes and further highlighted that over-expression of the two genes translates into significantly elevated indole-3-acetic acid levels. The described accumulation of YUC8 and YUC9 transcripts appears to be dependent on the canonical jasmonate receptor COI1, because the induction observed in wild type is completely missing in the coi1 genetic background (Fig. 2). Hence, our results point towards a novel connection through which jasmonate signaling and auxin biosynthesis are linked. Particularly interesting is the fact that jasmonate produced in response to mechanical wounding of leaves is sufficient to induce YUC9 expression in aerial tissue (Fig. 3).
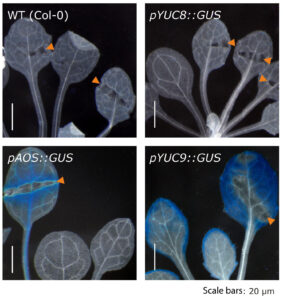
Figure 3. The wound-induced formation of methyl jasmonate in leaves is sufficient to trigger YUC9, but not YUC8 gene expression in leaves.
At the moment, our lab is trying to unravel both the molecular mechanism that mediate jasmonate-auxin crosstalk and the function of this relationship in physiological terms. Currently, we are undertaking microarray assays and directed plant hormone quantification experiments to verify our actual working-hypothesis that wounding initiates a signaling cascade, including the crosstalk between jasmonate, auxin, and ethylene, to answer mechanical stress or wounding by, e.g., blocking cell elongation and enforcing secondary cell wall growth (Fig. 4).